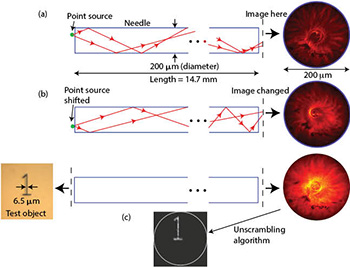
 (a) A point source is mapped onto an “image.” (b) The image changes when the point source is displaced. (c) A test object back-illuminated with an LED forms a complex image that can be unscrambled. All results are experimental.
(a) A point source is mapped onto an “image.” (b) The image changes when the point source is displaced. (c) A test object back-illuminated with an LED forms a complex image that can be unscrambled. All results are experimental.
High-resolution microscopy is essential for a wide range of scientific and engineering disciplines. However, microscopy in hard-to-reach parts of the body, such as the deep brain, is extremely challenging. Multiphoton microscopy is useful to image at depths of about 1 mm, but not much deeper.1 It requires high-power lasers, and achievable resolution is limited due to the long wavelengths. An alternative approach is to use miniaturized microscopes, but these require complex optical and electronic subsystems and are not readily capable of deep brain imaging.2 Recently, we applied a new computational technique to convert a simple needle into a high-resolution microscope.3
In a conventional imaging system, each point in an image corresponds to a unique point in the object, i.e., there is a one-to-one map between the object and the image. However, this is not strictly necessary. Each point on an object can be mapped onto many points on the image and captured on a sensor. As long as the intensity distribution formed on the sensor is unique for each object point and can be well-characterized, we can then apply computation to recover the object information. In a needle, we achieve this using multiple reflections of rays as they pass from a point source on one end to the other. If the point is slightly displaced, the intensity pattern at the other end of the needle changes. As a first step, we calibrate the intensity pattern at the output of the needle as a function of the position of the point source at the input. Since any incoherent object is a linear combination of point sources, one can apply a variety of algorithms to extract the object information from the scrambled image. We also showed that 3-D imaging in a volume in the vicinity of the needle is feasible as well.
We demonstrated this idea using an off-the-shelf needle and achieved a transverse resolution of approximately 1 μm. Since only a single frame is required to form the image, extremely fast microscopy becomes possible. We have also demonstrated the working principle using a variety of fluorescently tagged biological samples. If combined with photoswitchable fluorophores, this technique should enable a new mode of super-resolution imaging as well.
In summary, we used computational techniques to turn a needle into a fast, high-resolution microscope to enable deep brain imaging. Due to its simplicity and compact size, we believe this technology can have a significant impact on a variety of fields.
Researchers
Ganghun Kim and Rajesh Menon, University of Utah, USA
References
1. N.G. Horton et al. Nat. Photon. 7, 205 (2013).
2. K.K. Ghosh et al. Nat. Methods 8, 871 (2011).
3. G. Kim and R. Menon. Appl. Phys. Lett. 105, 061114 (2014).
